Our Services
VANGARD provides standard analysis for RNA-Seq, DNA-Seq, ChIP-Seq, and GRO/PRO-Seq, which implements "best practices" pipeline for the sequencing data provided.
ChIP-Seq Analysis
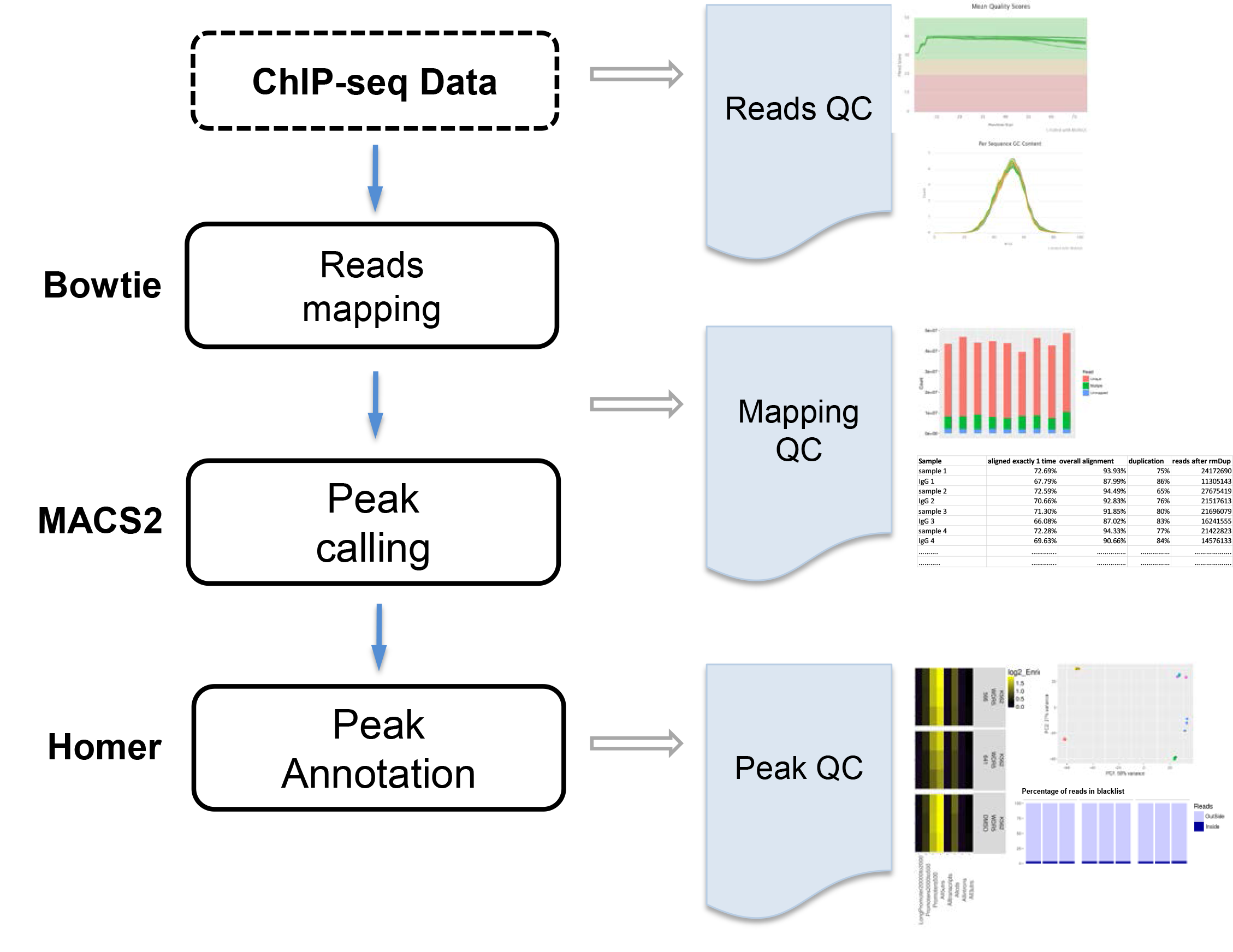
The goal is to identify protein binding sites. ChIP-seq data are mapped to the reference genome using Bowtie1, peaks are called using MACS22, and peaks are annotated by Homer. VANGARD will perform quality check, including raw reads quality (reads QC), mapping quality (mapping QC) and peak quality (peak QC) to ensure high quality results.
References
1. Langmead, B., Trapnell, C., Pop, M. and Salzberg, S.L. (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol, 10, R25.
2. Zhang, Y., Liu, T., Meyer, C.A., Eeckhoute, J., Johnson, D.S., Bernstein, B.E., Nusbaum, C., Myers, R.M., Brown, M., Li, W. et al. (2008) Model-based analysis of ChIP-Seq (MACS). Genome Biol, 9, R137.